A longer explanation:
I’m making a model for intervals between events. The data (timestamps of the events) for my model were collected with the timestamps having a resolution of one second (rounded down). The way these timestamps were collected wasn’t the most accurate, but I’m fairly certain that they’re usually off by one second at most. Hence I have a discrete uniform error distribution with support {-1,0,1} for the timestamps plus a maximum of one second of rounding error.
Since I’m modelling the intervals, the error distribution ends up being (after assuming independent errors for the timestamps and doing some simple combinatorics)
p(x) = \begin{cases}
1/9, &-2 \leq x < -1\\
2/9, &-1 \leq x < 0\\
3/9, &0 \leq x < 1\\
2/9, &1 \leq x < 2\\
1/9, &2 \leq x < 3\\
\end{cases}
I called this “podium” in the code since its shape resembles the shape of a winners’ podium. This might not be the “true” error distribution but the inference results seem to be somewhat unstable w.r.t. the choice of error distribution and I’d like to also see the results from this one.
For the underlying distribution of the unrounded intervals, I used Gamma. This also might not be the final choice but it felt like an OK first guess. It also has a nice feature of highlighting the issues with using the wrong error distribution in the model. I’m using mean/variance parametrization and the mean is estimated well from simulated data by just taking the rounding into account. The variance however need an correctly defined error distribution.
I felt that the most straightforward way for me to implement this error distribution with pymc was by constructing it as a mixture three different components as seen in the first post. I also tried the perhaps more readable way of using pm.Mixture but it wasn’t any faster.
I made the following toy example for testing the model
import numpy as np
import pymc as pm
from pymc.math import floor, log, exp
import pytensor.tensor as pt
import arviz
from pandas import DataFrame
def _create_data(data_count):
model = pm.Model()
with model:
raw = pm.Gamma(
"raw", mu=11, sigma=2 # mu set high and sigma low to avoid values < 3
)
rounded = pm.Deterministic("rounded", floor(raw))
uniform_error = pm.DiscreteUniform("uniform_error", lower=-1, upper=1)
pm.Deterministic("uniform", rounded + uniform_error)
podium_error = pm.Mixture(
"podium_error",
[5 / 9, 3 / 9, 1 / 9],
[
pm.DiscreteUniform.dist(-2, 2),
pm.DiscreteUniform.dist(-1, 1),
pm.DiscreteUniform.dist(0, 0),
],
)
pm.Deterministic("podium", rounded + podium_error)
data = pm.sample_prior_predictive(samples=data_count, random_seed=1).prior
return (
data.raw.values.flatten(),
data.rounded.values.flatten(),
data.uniform.values.flatten(),
data.podium.values.flatten(),
)
def _floor_dist(mu, sigma, size):
raw_dist = pm.Gamma.dist(mu=mu, sigma=sigma, size=size)
return pt.floor(raw_dist)
def _flexible_floor_logp(value, mu, sigma, lower, upper):
dist = pm.Gamma.dist(mu=mu, sigma=sigma)
density = exp(pm.logcdf(dist, value + upper)) - exp(pm.logcdf(dist, value + lower))
return log(density)
def _podium_logp(value, mu, sigma):
dist = pm.Gamma.dist(mu=mu, sigma=sigma)
density1 = exp(pm.logcdf(dist, value + 3)) - exp(pm.logcdf(dist, value - 2))
density2 = exp(pm.logcdf(dist, value + 2)) - exp(pm.logcdf(dist, value - 1))
density3 = exp(pm.logcdf(dist, value + 1)) - exp(pm.logcdf(dist, value))
return log(5 / 9 * density1 + 3 / 9 * density2 + 1 / 9 * density3)
def _create_models(data):
naive_model = pm.Model()
with naive_model:
mu = pm.Normal("mu", mu=0, sigma=10)
sigma = pm.HalfNormal("sigma", sigma=10)
mutable_data = pm.MutableData("data", data)
pm.Gamma("y", mu=mu, sigma=sigma, observed=mutable_data)
rounded_model = pm.Model()
with rounded_model:
mu = pm.Normal("mu", mu=0, sigma=10)
sigma = pm.HalfNormal("sigma", sigma=10)
mutable_data = pm.MutableData("data", data)
pm.CustomDist(
"y",
mu,
sigma,
dist=_floor_dist,
observed=mutable_data,
)
uniform_model = pm.Model()
with uniform_model:
mu = pm.Normal("mu", mu=0, sigma=10)
sigma = pm.HalfNormal("sigma", sigma=10)
mutable_data = pm.MutableData("data", data)
pm.CustomDist(
"y",
mu,
sigma,
-1,
2,
logp=_flexible_floor_logp,
observed=mutable_data,
)
podium_model = pm.Model()
with podium_model:
mu = pm.Normal("mu", mu=0, sigma=10)
sigma = pm.HalfNormal("sigma", sigma=10)
mutable_data = pm.MutableData("data", data)
pm.CustomDist(
"y",
mu,
sigma,
logp=_podium_logp,
observed=mutable_data,)
return naive_model, rounded_model, uniform_model, podium_model
def _test_model(model, data, rounded_data, uniform_data, podium_data):
summaries = {
"Correct": [11, 2],
}
with model:
if data is not None:
pm.set_data({"data": data})
samples = pm.sample(
nuts_sampler="blackjax", draws=100, chains=1, random_seed=1
)
summaries["Raw"] = arviz.summary(samples)["mean"]
if rounded_data is not None:
pm.set_data({"data": rounded_data})
samples = pm.sample(
nuts_sampler="blackjax", draws=100, chains=1, random_seed=1
)
summaries["Rounded"] = arviz.summary(samples)["mean"]
if uniform_data is not None:
pm.set_data({"data": uniform_data})
samples = pm.sample(
nuts_sampler="blackjax", draws=100, chains=1, random_seed=1
)
summaries["Uniform"] = arviz.summary(samples)["mean"]
if podium_data is not None:
pm.set_data({"data": podium_data})
samples = pm.sample(
nuts_sampler="blackjax",
draws=100,
chains=1,
random_seed=1,
initvals={"mu": 0, "sigma_log__": np.log10(11)},
)
summaries["Podium"] = arviz.summary(samples)["mean"]
summary = DataFrame(data=summaries)
print(summary)
def main():
data, rounded_data, uniform_data, podium_data = _create_data(10000)
naive_model, rounded_model, uniform_model, podium_model = _create_models(data)
_test_model(naive_model, data, rounded_data, uniform_data, podium_data)
_test_model(rounded_model, None, rounded_data, uniform_data, podium_data)
_test_model(uniform_model, None, None, uniform_data, podium_data)
_test_model(podium_model, None, None, None, podium_data)
if __name__ == "__main__":
main()
I compared the printed runtimes (e.g. Sampling time = 0:05:54) with the different models and data. For naive_model the runtimes were 6 seconds, for rounded_model and uniform_model roughly 120 seconds and for podium_model roughly 360 seconds.
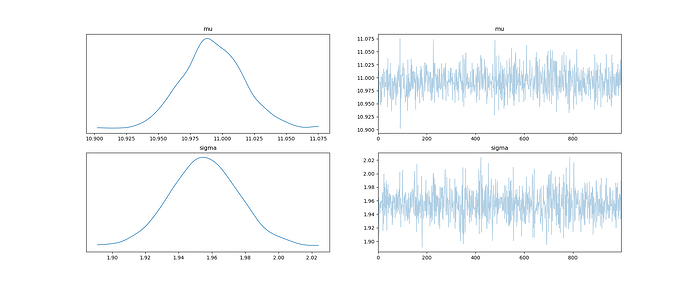
I also compared the sample means for mu and sigma. As expected, the estimated mu was off by 0.5 if the data was rounded and model didn’t account for this. If the model wasn’t specified correctly, estimated sigma was off by 0.18 - 0.36 depending on the model and data.