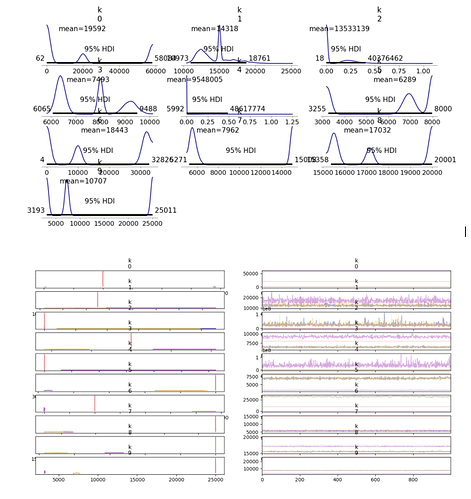
Hello all,
I have an issue in the speed of sampling. The code itself originally required performing multiple matrix inversions. Reading threads elsewhere, I saw that this was a potential avenue for bottlenecks, I therefore made the following adjustments.
- For smaller matrices, work the inverse out with a pen and paper, then code this directly.
- For larger matrices, use
pt.linalg.solveto obtain A^{-1} - Avoid use of loops, scans. Vectorise as much as possible.
In using the above, I was able to reduce NUTS-based sampling time down from 38 hours to around 3. This was for 1000 samples and 1000 tuning steps, so not much in terms of sampling demand.
Despite the speed, inference is running correctly, so my question really is one of speed. Is circa 3-hours as good as it gets? Sampling seems to start off quick enough but grinds to a very slow rate.
Here is a section of code which causes the issue, happy to provide more if needed. The general steps are as follows:
-
Construct a series of (\omega x 2 x 2) matrices using priors (a third dimension, \omega, handles the fact that 2 x 2 matrices vary with frequency).
-
Arrange into a block diagonal format. This is
Y_combinedin the code snippet. -
Compose a further block diagonal with a variable defined outside of the model context (
Y_shared). In the code, I provide 2 options for this: Option 1 uses a preallocated zero tensor which is part-filled withY_shared. I then usept.set_subtensorto add in the remaining block diagonal matrixY_combined, within the model context as shown in the snippet.
Y_{full} = \left[ \begin{array}{c|c} \mathbf{Y_{shared}} & 0 \\ \hline 0 & \mathbf{Y_{combined} } \end{array} \right]
For Option 2 (commented out in the snip below), I simply use the block diagonal function (pytensor.tensor.linalg import block_diag as block_diag_pt). Note that this option seems to restrict the use of gradient-based sampling and I don’t yet have a work around.
-
Perform some calculations, using
pt.linalg.solvewhere an inverse is needed, as described previously. -
Sample.
Any pointers would be hugely appreciated.
with pm.Model() as model:
# Priors
k = pm.LogNormal("k", mu=np.log(10000), sigma=1, shape=(10,))
eta = pm.Beta("eta", alpha=2, beta=2, shape=(10,))
sigma = pm.HalfNormal("sigma", 0.1)
# Deterministic transformations
k_vals = pm.Deterministic("k_vals", k)
eta_vals = pm.Deterministic("eta_vals", eta)
# Manually construct inverse for smaller matrices
det_Z = 1 / ((k * winv_pt[:, None])**2 * (1 + eta**2))
Y11 = (k * eta * winv_pt[:, None]) * det_Z
Y12 = (-k * winv_pt[:, None]) * det_Z
Y21 = (k * winv_pt[:, None]) * det_Z
Y22 = (k * eta * winv_pt[:, None]) * det_Z
# Construct 2x2 matrices/tensors
upper_stack = pt.stack([Y11, Y12], axis=2) # shape (len(w), 1, 2)
lower_stack = pt.stack([Y21, Y22], axis=2) # shape (len(w), 1, 2)
Y_combined = pt.zeros((len(f), 2*num_coupled, 2*num_coupled))
# Fill each 2x2 block into the correct diagonal position
for i in range(10): # there are 10 2x2 matrices to be arranged in block diagonal fashion
idx = 2 * i
Y_isol_matrix_pt = pt.set_subtensor(
Y_isol_matrix_pt[:, idx:idx+2, idx:idx+2],
pt.stack([
pt.stack([Y11[:, i], Y12[:, i]], axis=1),
pt.stack([Y21[:, i], Y22[:, i]], axis=1)
], axis=1)
)
# Option 1 - preallocate a matrix/tensor with zeros and use set_subtensor to fill in entries.
# isolator_start_index = Y_mat_shared.eval().shape[1]
Y_full = pt.set_subtensor(Y_full[:, isolator_start_index:isolator_start_index+2*num_coupled, isolator_start_index:isolator_start_index+2*num_coupled], Y_combined )
# Option 2 - using block diag to construct directly
# Y_full = block_diag_pt(Y_mat_shared2,Y_isol_matrix_pt)
# Compute the bracket and inverse
bracket_pt1 = B_shared @ Y_full @ Bt_shared
bracket_pt = pt.linalg.solve(bracket_pt1, pt.eye(bracket_pt1.shape[1]))
# Compute Yc_pt
Yc_pt = Y_full - Y_full @ Bt_shared @ bracket_pt @ B_shared @ Y_full
likelihood = pm.Normal("likelihood", mu=pt.flatten(Yc_pt), sigma=sigma, observed=data)
# Sampling
trace = pm.sample(1000,
tune=1000,
chains=4,
cores=15,
nuts_sampler='pymc',
nuts={"max_treedepth": 11},
return_inferencedata=True,
discard_tuned_samples=False,
random_seed=42)