I might be losing my mind here, would definitely appreciate input from a kind soul.
I think I’ve misunderstood xarray indexing, but there could be something else going on too. This question has become apparent during my plotting model posteriors.
Background
I have a model that uses an LKJCholeskyCov to create an MvNormal. I name the dims, and I use the unpacked compute_corr=True parameter to leave me a useful _corr in the trace.
Relevant code fragment:
self.coords.update({'b0_names': ['gc_omega_b0', 'gc_theta_b0', 'ulr_b0']})
...
sd_dist = pm.InverseGamma.dist(alpha=11, beta=10)
chol, corr_, stds_ = pm.LKJCholeskyCov('lkjcc', n=3, eta=2,
sd_dist=sd_dist, compute_corr=True)
b0 = pm.MvNormal('b0', mu=0, chol=chol, dims='b0_names')
After sampling I can plot posteriors for b0:
axs_pp = az.plot_pair(mdlb6.idata, var_names=rvs, divergences=True,
marginals=True, figsize=(5, 5))
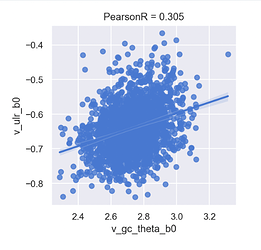
Observe:
- There appears to be positive correlation between
ulr_b0andgc_theta_b0(the others are negative), and in conventional lower triangle array indexing, this is plotted at index 4:
[[0 - -]
[1 2 -]
[3 4 5]]
- If I want the 2D
i, jindex for this I can usenp.tril_indicesandb0_namesand recreate the axis labels shown on the plot. Just to confirm, index 4 is wherei=2,j=1, and the relevant pair ofb0_namesis('ulr_b0', 'gc_theta_b0'), and the plot showsxlabel='b0\ngc_theta_b0'
i, j = np.tril_indices(n=3, k=-1)
print(i, j)
> [1 2 2] [0 1 1]
[(coords['b0_names'][a], coords['b0_names'][b]) for a, b in zip(i, j)]
> [('gc_theta_b0', 'gc_omega_b0'),
> ('ulr_b0', 'gc_omega_b0'),
> ('ulr_b0', 'gc_theta_b0')]
axs_pp[2][1]
> <AxesSubplot:xlabel='b0\ngc_theta_b0'>
-
Finally to note if I read along
i, jI can say the order of paired correlations is[negative, negative, positive] -
All good so far
My surprise observation
If I now use i, j to index lkj_corr and then plot_posterior, I get plots that appear to show a reversed order [positive, negative, negative]:
axs = az.plot_posterior(
mdlb6.idata.posterior['lkjcc_corr'].values[:, :, i, j],
ref_val=[0, 0, 0], figsize=(12, 2))
… I can plot this in my expected order if I reverse i and j and then index lkjcc_corr
axs = az.plot_posterior(
mdlb6.idata.posterior['lkjcc_corr'].values[:, :, i[::-1], j[::-1]],
ref_val=[0, 0, 0], figsize=(12, 2))
Just to isolate this further, let’s view index 4 as above where i=2, j=1.. yep that’s definitely not what I expected
axs = az.plot_posterior(
mdlb6.idata.posterior['lkjcc_corr'].values[:, :, 2, 1],
ref_val=[0, 0, 0], figsize=(8, 2))
If I view the xarray in Notebook, the dimensions look reasonable to me and the lowest 2D unit of dimensions is 3x3, has an identity diagonal - it looks as I would expect. Obviously this means that merely swapping i, j would get the same mirrored value from the upper triangle, so the order doesn’t matter
Even more concretely
It seems that when I try to index so as to get cell 4, I instead get cell 1, which is a “flip and rotate” or a mirror around the anti-diagonal
e.g.
a = np.array([
[0, np.nan, np.nan],
[1, 2, np.nan],
[3, 4, 5],
])
print(a)
i, j = np.tril_indices(n=3, k=-1)
print(i, j)
print(a[i[-1:], j[-1:]])
print(b)
b = np.rot90(np.fliplr(a), k=3)
print(b[i[-1:], j[-1:]])
>>
[[ 0. nan nan]
[ 1. 2. nan]
[ 3. 4. 5.]]
[1 2 2] [0 0 1]
[4.]
[[ 5. nan nan]
[ 4. 2. nan]
[ 3. 1. 0.]]
[1.]
My stupid question
Does lkjcc_corr somehow have an inverted index?
Or: what have I misunderstood about xarray indexing?
Many thanks if you made it this far!
Relevant env details:
pymc3=3.11.4
arviz=0.12.1