Hello,
I am trying to sample from a Gaussian Mixture (pm.NormalMixture()) that has two (as an illustratory example, in practice there’s more, e.g. 8 - 36) Normals mixed, although their means are too far away from each other.
Here: N_1(300, 100^2) and N_2(70000, 5000^2)
The mixture is mixed with weights 0.7 \cdot N_1 + 0.3 \cdot N_2, where the weights are based on a Dirichlet distribution, which is above the NormalMixture in the hierarchy.
Although when I sample the posterior of the NormalMixture, there is mainly the N_2, which should not happen. It should be more of the N_1.
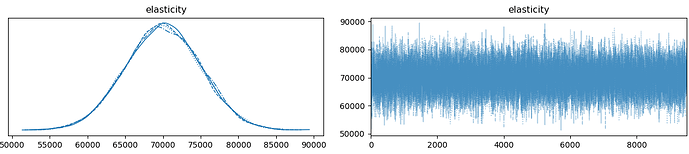
The code used to generate the following graphs:
with pm.Model() as miniBN:
mat = pm.Dirichlet('mat', a=mat_probs)
elasticity = pm.NormalMixture('elasticity', w=mat, mu=ELAS_MUS, sigma=ELAS_SIGS)
burn_in = 500
trace = pm.sample(10000, tune=5000, target_accept=0.9)
chain = trace[burn_in:]
pm.plot_trace(chain)
pm.plot_posterior(chain)
plt.show()
Question:
Am I just undersampling? Should I sample more? It does not seem to help if I ramp up the sampling to say pm.sample(50000, tune=1000, target_accept=0.9). What is the proper way of handling such situations?