I’m not sure I understand the question correctly, please correct me if I misunderstood something.
You want to compute the gradient a posterior estimate of a parameter with respect to the x-positions where a gp was evaluated?
So for instance we have the following gp:
import pymc as pm
import pytensor.tensor as pt
import numpy as np
import matplotlib.pyplot as plt
np.random.seed(2)
n = 10
X = np.linspace(0, 10, n)
ell_true = 5.0
eta_true = 8.0
cov_func = eta_true**2 * pm.gp.cov.Matern52(1, ell_true)
mean_func = pm.gp.mean.Zero()
f_true = np.random.multivariate_normal(
mean_func(X[:, None]).eval(), cov_func(X[:, None]).eval() + 1e-8 * np.eye(n), 1
).flatten()
sigma_true = 0.02
y = f_true + sigma_true * np.random.randn(n)
plt.plot(X, y)
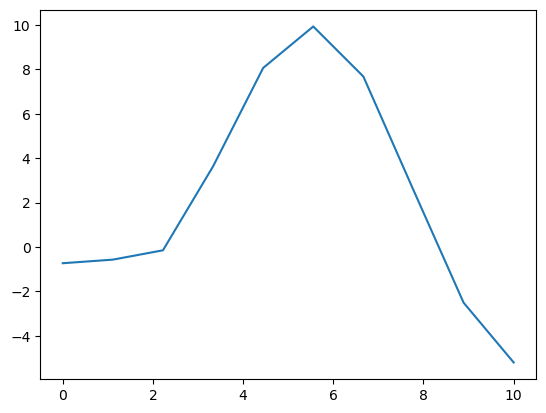
We can then model this to get a posterior over the mean of the gp:
with pm.Model(coords={"x_dim": X}) as model:
ell = pm.InverseGamma("ell", mu=1.0, sigma=0.6)
eta = pm.Exponential("eta", scale=3.0)
cov = eta**2 * pm.gp.cov.Matern52(1, ls=ell)
gp = pm.gp.Marginal(cov_func=cov)
sigma = pm.HalfCauchy("sigma", beta=5)
X = pm.Data("X_data", X)
y = pm.Data("y_data", y)
y_ = gp.marginal_likelihood("y", X=X[:, None], y=y, sigma=sigma)
mu, var = gp._predict_at(
X[:, None], diag=True
)
sd = np.sqrt(var)
gp_mu = pm.Deterministic("gp_mu", mu.mean())
And you would now like to know what the gradient of gp_mu.mean(["draw", "chain"]) is with respect to X?
We can use a nice property of the posterior distribution. If we have any well behaved function f(\theta, \eta) where \theta are the parameters of the model and \eta are any other hyperparameters, then we can compute derivatives of F(\eta) = \int P(\theta|y, \eta) f(\theta, \eta) d\theta with
\frac{dF}{d\eta} = \int P(\theta|y, \eta) \left[\frac{df(\theta, \eta)}{d\eta} + (f(\theta, \eta) - F(\eta)) \frac{d}{d\eta}\log(P(y|\theta, \eta)) \right]
And given samples from P(\theta|y, \eta) we can approximate both integrals using their monte-carlo estimate (ie just the mean over the draws).
In this case we want \eta = X and f(\theta, \eta) = \text{gp_mu}, so that F(\eta) is just the posterior mean of gp_mu.
We can then compute the gradient as follows (adding to the model above):
with pm.Model(coords={"x_dim": X}) as model:
logp = model.logp()
# The derivatives in the integral above
pm.Deterministic("dlogp_dx", pt.grad(logp, X), dims=("x_dim",))
pm.Deterministic("dmu_dx", pt.grad(gp_mu, X), dims=("x_dim",))
tr = pm.sample(nuts_sampler="numpyro")
And to get the gradient:
f = tr.posterior.gp_mu
F = f.mean(["draw", "chain"])
(tr.posterior.dmu_dx + (f - F) * tr.posterior.dlogp_dx).mean(["draw", "chain"]).plot()

Which I think makes sense? For instance if the first or second x-value was a bit smaller or bigger, our estimate for the mean wouldn’t change much. But if the third x-value was bigger, then the peak would start later, and be therefore smaller, meaning the mean of the GP should be smaller.
If you don’t want to compute the full posterior but only the MAP, you can use the implicit function theorem to get the derivatives of the MAP-estimates. The easiest way to do that in practice is probably to export the model to jax, and then use optjax or optimistix to compute the MAP, those implement those derivatives already.
I hope this wasn’t too much in one go with too little explanation, feel free to ask if something doesn’t make sense to you.


