Hi, I have a question regarding a relatively simply probabilistic programming example that involves discrete variables. The example is taken from Chapter 1 of the book “Practical Probabilistic Programming” (see Table 1.1 here: Chapter 1. Probabilistic programming in a nutshell - Practical Probabilistic Programming (manning.com)).
I implemented this scenario in WebPPL ( WebPPL - probabilistic programming for the web) (see this gist: webppl example (github.com)) and now I am wondering whether it is possible to implement the same example in pymc3.
Here is the scenario:
- Assume the weather can have two states
sunnyandnot sunnywith the following (prior) probabilities:- Sunny 0.2
- Not sunny 0.8
- Based on the weather, we will be greeted with different greetings according to the following probabilistic logic:
- If today’s weather is sunny
- “Hello, world!” 0.6
- "Howdy, universe!” 0.4
- If today’s weather isn’t sunny
- “Hello, world!” 0.2
- "Oh no, not again” 0.8
- If today’s weather is sunny
No, we want to do two things:
- Simulate today’s greeting based on the (prior) probability of seeing sunny weather
- Infer today’s weather based on an observed greeting
I implemented the first task using pm.sample_prior_predictive() with this code:
import altair as alt
import numpy as np
import pandas as pd
import pymc as pm
from aesara.ifelse import ifelse
g0 = "Hello, World!"
g1 = "Howdy, universe!"
g2 = "Oh no, not again!"
greetings = np.array([g0, g1, g2])
weather_states = np.array(["cloudy", "sunny"])
def greeting(model, sunny):
if model is None:
model = pm.Model()
with model:
p_choice = ifelse(sunny, 0.6, 0.2)
altAnswer = pm.Deterministic('altAnswer', ifelse(sunny, 1, 2))
greeting = pm.Deterministic('GreetingToday', ifelse(pm.Bernoulli("GreetingChoice", p_choice), 0, altAnswer))
return model
def predictGreetingToday():
p_sunny = 0.2
with pm.Model() as model:
sunny = pm.Bernoulli("sunny", p_sunny)
greeting(model, sunny)
return model
trace = pm.sample_prior_predictive(1000, model=predictGreetingToday())
# Plot the results
sunny_df = pd.DataFrame(
{"weather": weather_states[np.array(trace.prior["sunny"]).reshape(-1)]}
)
weather_chart = alt.Chart(
sunny_df.value_counts(normalize=True).reset_index().rename(columns={0: "count"})
).mark_bar().encode(x="weather:N", y="count:Q")
greeting_df = pd.DataFrame(
{"greeting": greetings[np.array(trace.prior["GreetingToday"]).reshape(-1)]}
)
greeting_chart = alt.Chart(
greeting_df.value_counts(normalize=True).reset_index().rename(columns={0: "count"})
).mark_bar().encode(x="greeting:N", y="count:Q")
weather_chart | greeting_chart
giving something like this:
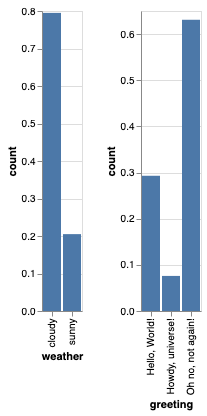
However, I am struggling to implement the 2nd task, i.e., to do some inference. The main problem is that it is not possible to do inference on the deterministic variable greeting, which is what we observe. I think I could make it work implementing the backward logic manually, i.e., do something like this:
if greeting == g2:
altAnswer_obs = g2
sunny = False
elif greeting == g1:
altAnswer_obs = g1
sunny = True
else:
...
# not sure what do do here
However, even if that would work, this is not really the point, because the point of Probabilistic Programming, generally speaking, is to do this kind of probabilistic reasoning on its own, right?
I understand that this is not really the typical use case of pymc3 but I am trying to understand the difference between different PPLs, like e.g. webPPL and pymc3, so is it possible at all do implement this scenario in pymc3?
WebPPL can solve this task either with exact enumeration (which is feasible for these simple cases with discrete variables) or rejection sampling.
If not, are there other PPLs implemented in python that can do this?
