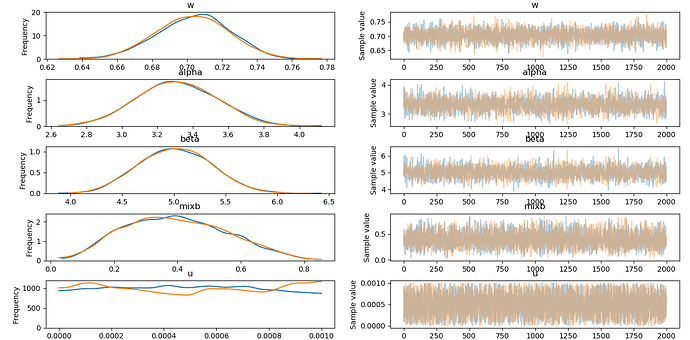
This is fantastic and is much appreciated! Thank you!
I have been trying to recreate the above model construction using my own construct_model function that makes use of a design matrix of shape (M, N) and observed data of shape (M, 1). Unfortunately, naively taking the above code and inserting the weights and mixture distribution results in the following error:
---------------------------------------------------------------------------
AttributeError Traceback (most recent call last)
/anaconda3/envs/audience-concentration-forecast-service-env/lib/python3.6/site-packages/pymc3/distributions/mixture.py in _comp_means(self)
119 try:
--> 120 return tt.as_tensor_variable(self.comp_dists.mean)
121 except AttributeError:
AttributeError: 'list' object has no attribute 'mean'
During handling of the above exception, another exception occurred:
IndexError Traceback (most recent call last)
<ipython-input-18-69ac3b7b6842> in <module>
----> 1 model = construct_model(data, preprocessors)
<ipython-input-17-51f9055e6674> in construct_model(data, preprocessors, fit_intercept)
63 w=[weights, 1 - weights],
64 comp_dists=[measured.distribution, unmeasured.distribution],
---> 65 observed=success_rate)
66
67 return model
/anaconda3/envs/audience-concentration-forecast-service-env/lib/python3.6/site-packages/pymc3/distributions/distribution.py in __new__(cls, name, *args, **kwargs)
39 raise TypeError("observed needs to be data but got: {}".format(type(data)))
40 total_size = kwargs.pop('total_size', None)
---> 41 dist = cls.dist(*args, **kwargs)
42 return model.Var(name, dist, data, total_size)
43 else:
/anaconda3/envs/audience-concentration-forecast-service-env/lib/python3.6/site-packages/pymc3/distributions/distribution.py in dist(cls, *args, **kwargs)
50 def dist(cls, *args, **kwargs):
51 dist = object.__new__(cls)
---> 52 dist.__init__(*args, **kwargs)
53 return dist
54
/anaconda3/envs/audience-concentration-forecast-service-env/lib/python3.6/site-packages/pymc3/distributions/mixture.py in __init__(self, w, comp_dists, *args, **kwargs)
84
85 try:
---> 86 self.mean = (w * self._comp_means()).sum(axis=-1)
87
88 if 'mean' not in defaults:
/anaconda3/envs/audience-concentration-forecast-service-env/lib/python3.6/site-packages/pymc3/distributions/mixture.py in _comp_means(self)
122 return tt.squeeze(tt.stack([comp_dist.mean
123 for comp_dist in self.comp_dists],
--> 124 axis=1))
125
126 def _comp_modes(self):
/anaconda3/envs/audience-concentration-forecast-service-env/lib/python3.6/site-packages/theano/tensor/basic.py in stack(*tensors, **kwargs)
4707 dtype = scal.upcast(*[i.dtype for i in tensors])
4708 return theano.tensor.opt.MakeVector(dtype)(*tensors)
-> 4709 return join(axis, *[shape_padaxis(t, axis) for t in tensors])
4710
4711
/anaconda3/envs/audience-concentration-forecast-service-env/lib/python3.6/site-packages/theano/tensor/basic.py in <listcomp>(.0)
4707 dtype = scal.upcast(*[i.dtype for i in tensors])
4708 return theano.tensor.opt.MakeVector(dtype)(*tensors)
-> 4709 return join(axis, *[shape_padaxis(t, axis) for t in tensors])
4710
4711
/anaconda3/envs/audience-concentration-forecast-service-env/lib/python3.6/site-packages/theano/tensor/basic.py in shape_padaxis(t, axis)
4599 if not -ndim <= axis < ndim:
4600 msg = 'axis {0} is out of bounds [-{1}, {1})'.format(axis, ndim)
-> 4601 raise IndexError(msg)
4602 if axis < 0:
4603 axis += ndim
IndexError: axis 1 is out of bounds [-1, 1)
The snippet below abstractly constructs the model:
import logging
import numpy as np
import pymc3 as pm
import scipy.sparse
import theano
def assemble_coefficient_vector(feature_shapes, default_sd=2.5):
"""Using the dictionary of features and their dimensions (shapes), assemble
the coefficient vector to be used for regression. We assume that the design_matrix
does not contain a vector of 1's for the intercept.
"""
coefficient_vector = []
for feature in feature_shapes:
# Get number of features
N = feature_shapes[feature]
sigma = pm.HalfStudentT(f'σ_{feature}', nu=4, sd=default_sd)
coefficients = pm.StudentT(f'β_{feature}', nu=4, mu=0, sd=sigma, shape=(N, 1))
coefficient_vector.append(coefficients)
coefficient_vector = theano.tensor.concatenate(coefficient_vector, axis=0)
return coefficient_vector
def construct_model(data, preprocessors):
# create model just relies on preprocessors (design_matrix, response) and data
design_matrix_preprocessor = preprocessors['design_matrix']
response_preprocessor = preprocessors['response']
feature_shapes = mp.get_feature_shapes(design_matrix_preprocessor)
design_matrix = design_matrix_preprocessor.transform(data)
M, N = design_matrix.shape
response = response_preprocessor.transform(data)
X = theano.sparse.as_sparse_variable(design_matrix)
success_rate = theano.tensor.as_tensor_variable(response)
with pm.Model() as model:
# Of shape (N, 1)
coefficient_vector = assemble_coefficient_vector(feature_shapes, fit_intercept=fit_intercept)
# Multiply matrix M x N times vector N x 1 to get M x 1 vector
mu = theano.sparse.dot(X, coefficient_vector)
kappa = pm.HalfStudentT('κ', sd=2.5)
likelihood = pm.StudentT('likelihood', nu=4, mu=mu, sd=kappa, observed=success_rate)
return model
Is there something obviously wrong that I am doing?
And sorry, this is the construct_model definition with the modifications:
def construct_model(data, preprocessors, fit_intercept=False):
# create model just relies on preprocessors (design_matrix, response) and data
design_matrix_preprocessor = preprocessors['design_matrix']
response_preprocessor = preprocessors['response']
ZERO = float(response_preprocessor.transform(pd.DataFrame({'success_rate': [0.0]})))
ONE = float(response_preprocessor.transform(pd.DataFrame({'success_rate': [1.0]})))
feature_shapes = mp.get_feature_shapes(design_matrix_preprocessor)
design_matrix = design_matrix_preprocessor.transform(data)
M, N = design_matrix.shape
# Add a vector of ones to first column of design_matrix.
if fit_intercept:
intercept_design_matrix = np.concatenate([np.ones(shape=(M, 1)), design_matrix.todense()], axis=1)
design_matrix = scipy.sparse.csr_matrix(intercept_design_matrix)
response = response_preprocessor.transform(data)
X = theano.sparse.as_sparse_variable(design_matrix)
success_rate = theano.tensor.as_tensor_variable(response)
with pm.Model() as model:
weights = pm.Beta('w', 1, 1)
coefficient_vector = assemble_coefficient_vector(feature_shapes, fit_intercept=fit_intercept)
mu = theano.sparse.dot(X, coefficient_vector)
kappa = pm.HalfStudentT('κ', nu=4, sd=2.5)
measured = pm.StudentT('measured', nu=4, mu=mu, sd=kappa, shape=(M, 1))
unmeasured = pm.Normal('unmeasured', mu=ZERO, sd=1e-3, shape=(M, 1))
mixture = pm.Mixture('mixture',
w=[weights, 1 - weights],
comp_dists=[measured.distribution, unmeasured.distribution],
observed=success_rate)
return model