I get few problems about Pytensor. My major is physics, I use pymc and pytensor to process some signal.In my problem ,the signal’s expression F(t) can be divided as physical signal ,assume it as f(t) and the response of instrument, which is called IRF.The signal can be seen as the convolution of f(t) and IRF.there are some parameters in the F(t).I want use Bayes method to fit the curve, and find the parameters in it.
The first problem is the convolution in PyTensor.I’m not familiar with CNN or scan and I‘m just begin to learn it.I’m only familiar with how convolve works in numpy. I have asked here how convolve1d works in Pytensor, I’m wondering if it works the same as numpy or scipy ?I did‘t find the tutorial about 1d convolve in Pytensor.
The second problem is how to assign a certain part of a tensor to another tensor? If in numpy ,I use
import pytensor.tensor as pt
from pytensor.tensor import conv
import numpy as np
A=np.array([1,2,3,4,5,6,7])
B=np.array([1,2,3,4,5,6,7,0,0])
C=np.array([5,6,7])
D=np.convolve(A,C)
G=pt. fvector('G')
E=conv.causal_conv1d(A[None,None,:],C[None,None,:],filter_shape=(None,None,3),input_shape=(None,None,9)).squeeze()
F=conv.causal_conv1d(B[None,None,:],C[None,None,:],filter_shape=(None,None,3)).squeeze()
for i in range(4):
G[i] = E[i]
print(D)
print(E.eval())
print(F.eval())
to assign the first five elements of E to G, but if in PyTensor, G[i] = E[I] isn’t work ,what should I do?
My third problem is how to evaluate the result of sampling,
from pytensor.tensor import conv
def Ft(A,sigma2,mu2):
tau=1/sigma2**2
ft1=pt.zeros(num)
ft1=A*pm.math.sqrt(tau/(2*np.pi))*pm.math.exp(-tau/2*(t-mu2)**2)
IRF1=IRF[None,None,:]
ft2=ft1[None,None,:]
ft=(conv.causal_conv1d(ft2, IRF1, filter_shape=(1,1,IRF1.shape[2]))).squeeze()
return ft
#%%
with pm.Model() as final_model:
amp = pm.Uniform('amp',lower=-1.0,upper=0)
mu1 = pm.Uniform('mu1', lower=10,upper=20)
sigma1 = pm.Uniform('sigma1',lower=0,upper=5)
y_observed=pm.Normal(
"y_observed",
mu= Ft(amp,sigma1,mu1),
sigma=noise_sig,
observed=v4_s,
)
output = pm.Deterministic('output',Ft(amp,sigma1,mu1))
prior = pm.sample_prior_predictive()
posterior_f = pm.sample(draws =1000, target_accept = 0.9,chains=4,cores=4)
posterior_f = pm.sample_posterior_predictive(posterior_f, extend_inferencedata=True)
az.plot_trace(posterior_f, var_names = ['amp','mu1','sigma1'])
result=az.summary(posterior_f, var_names = ['amp','mu1','sigma1'])
print(result)
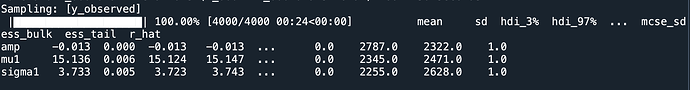
the results are:

seems the accuracy is ok,
I drew a graph based on the parameters deduced, and the results are as follows, compared to the original data, the error is very large, it seems like PyMC converged to the wrong result.

I’m a freshman user of Pytensor,so any advise is helpful.Thanks.