I have a hierarchical logistic regression model that aims to determine the probability of a success given a number of trials. The model itself is similar to the below Kruschke diagram with the difference being the different priors used:
For the priors, I’ve followed Gelman et. al’s advice on standardizing the covariates and placing weak priors around the coefficients, specifically t_4(mu=0, sd=1) priors. For the intercept I’m placing a t_4(mu=0, sd=5) prior and for the \kappa parameter I’m using an Exp(lam=1e-3) prior which is weak on the number of trials present in the data.
I’m wondering if the coefficient values that I’m receiving after inference are indicative of some problem with the model; The coefficient values on the logit scale are really wide and do not provide any distinction among them:
However, when performing posterior predictive checks, the model is providing reasonable estimates of the data. Should I be worried about these coefficient values even if the model is providing reasonable results?
Did you do all the diagnosis check? Divergence, rhat and effective sample size, energy plot, bfmi. Would be a good idea to check loo and the k hat from loo as well.
Yeah, from what I can tell there’s nothing out of the ordinary in terms of convergence. There were a total of less than 40 reported divergences out of the 2000 used in the final distribution, but that shouldn’t lead to extremely biased inference right?
Energy Error Distribution:

depth for both chains:

Step Size bar for both chains:

Mean tree accept histogram (mean was 0.8 which matches target_accept given on initialization)
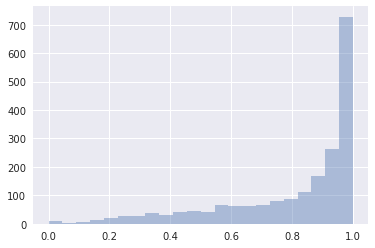
Energy vs energy diff

Further, the Rhat for all parameters was between 0.99 and 1.01 and the minimum effective sample size across all parameters was around 350. The BFMI was 0.9683649816013548, but I’m not sure what this parameter is (I’m assuming it should be near 1).
I’m not sure what the khat of the loo is, but here is the LOO output for the model/trace
LOO_r(LOO=121805.38669244738, LOO_se=626.8930392116976, p_LOO=11.25017988102627, shape_warn=0)
Do you agree with my conclusion about approximate convergence?
1 Like
Divergence, even just 1, could be an indication of bias estimation. While the rest of the diagnosis all look good, you should do a pairplot and check whether the divergence is systematic or not. If so, more informative prior usually helps; if not, increase the target accept and see if you can get rid of the divergences.
Hmm understood. If I am able to have a posterior with no divergences but the trace still looks as above, is there cause for concern then?
I’m rerunning with a target_accept = 0.9. Will report back with the results in 30 min.
Increasing the target_accept to 0.95 resulted in 2 divergences and the diagnosis remains the same. I’ll try a more informative prior to see if that changes things. So as long as the divergences are gone and the model reasonably mimics the data, these coefficient values should be fine?
1 Like
Just wanted to pipe in that this is an awesome job of convergence checking, and I am bookmarking for next time I have a difficult model.






